In Exercise 2.01, Building a Hierarchy, you implemented hierarchical clustering using what is known as Centroid Linkage. Linkage is the concept of determining how you can calculate the distances between clusters and is dependent on the type of problem you are facing. Centroid linkage was chosen for Exercise 2.02, Applying Linkage Criteria, as it essentially mirrors the new centroid search that we used in k-means. However, this is not the only option when it comes to clustering data points. Two other popular choices for determining distances between clusters are single linkage and complete linkage.
Single Linkage works by finding the minimum distance between a pair of points between two clusters as its criteria for linkage. Simply put, it essentially works by combining clusters based on the closest points between the two clusters. This is expressed mathematically as follows:
dist(a,b) = min( dist( a[i]), b[j] ) )
In the preceding code, a[i] is the ith point within first cluster where b[j] is jth point of second cluster.
Complete Linkage is the opposite of single linkage and it works by finding the maximum distance between a pair of points between two clusters as its criteria for linkage. Simply put, it works by combining clusters based on the furthest points between the two clusters. This is mathematically expressed as follows:
dist(a,b) = max( dist( a[i]), b[j] ) )
In the preceding code, a[i] and b[j] are ith and jth point of first and second cluster respectively. Determining what linkage criteria is best for your problem is as much art as it is science, and it is heavily dependent on your particular dataset. One reason to choose single linkage is if your data is similar in a nearest-neighbor sense; therefore, when there are differences, the data is extremely dissimilar. Since single linkage works by finding the closest points, it will not be affected by these distant outliers. However, as single linkage works by finding the smallest distance between a pair of points, it is quite prone to the noise distributed between the clusters. Conversely, complete linkage may be a better option if your data is distant in terms of inter-cluster state; complete linkage causes incorrect splitting when the spatial distribution of cluster is fairly imbalanced. Centroid linkage has similar benefits but falls apart if the data is very noisy and there are less clearly defined "centers" of clusters. Typically, the best approach is to try a few different linkage criteria options and see which fits your data in a way that's the most relevant to your goals.
Exercise 2.02: Applying Linkage Criteria
Recall the dummy data of the eight clusters that we generated in the previous exercise. In the real world, you may be given real data that resembles discrete Gaussian blobs in the same way. Imagine that the dummy data represents different groups of shoppers in a particular store. The store manager has asked you to analyze the shopper data in order to classify the customers into different groups so that they can tailor marketing materials to each group.
Using the data we generated in the previous exercise, or by generating new data, you are going to analyze which linkage types do the best job of grouping the customers into distinct clusters.
Once you have generated the data, view the documents supplied using SciPy to understand what linkage types are available in the linkage function. Then, evaluate the linkage types by applying them to your data. The linkage types you should test are shown in the following list:
['centroid', 'single', 'complete', 'average', 'weighted']
We haven't covered all of the previously mentioned linkage types yet – a key part of this activity is to learn how to parse the docstrings that are provided using packages to explore all of their capabilities. Follow these steps to complete this exercise:
- Visualize the
x dataset that we created in Exercise 2.01, Building a Hierarchy:from scipy.cluster.hierarchy import linkage, dendrogram, fcluster
from sklearn.datasets import make_blobs
import matplotlib.pyplot as plt
%matplotlib inline
- Generate a random cluster dataset to experiment on.
X = coordinate points, y = cluster labels (not needed):X, y = make_blobs(n_samples=1000, centers=8, \
n_features=2, random_state=800)
- Visualize the data, as follows:
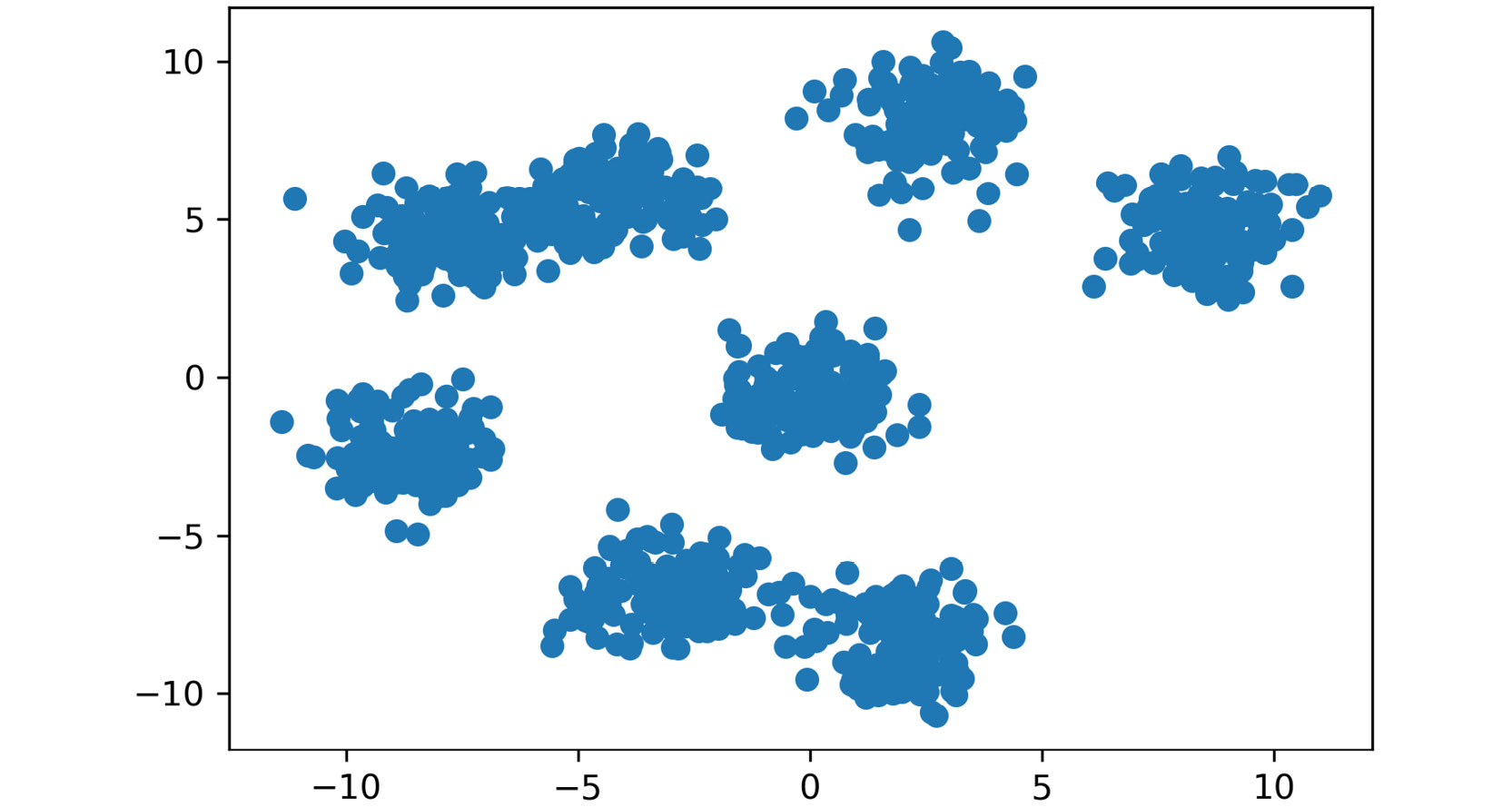
plt.scatter(X[:,0], X[:,1])
plt.show()
The output is as follows:
Figure 2.14: A scatter plot of the generated cluster dataset
- Create a list with all the possible linkage method hyperparameters:
methods = ['centroid', 'single', 'complete', \
'average', 'weighted']
- Loop through each of the methods in the list that you just created and display the effect that they have on the same dataset:
for method in methods:
distances = linkage(X, method=method, metric="euclidean")
clusters = fcluster(distances, 3, criterion="distance")
plt.title('linkage: ' + method)
plt.scatter(X[:,0], X[:,1], c=clusters, cmap='tab20b')
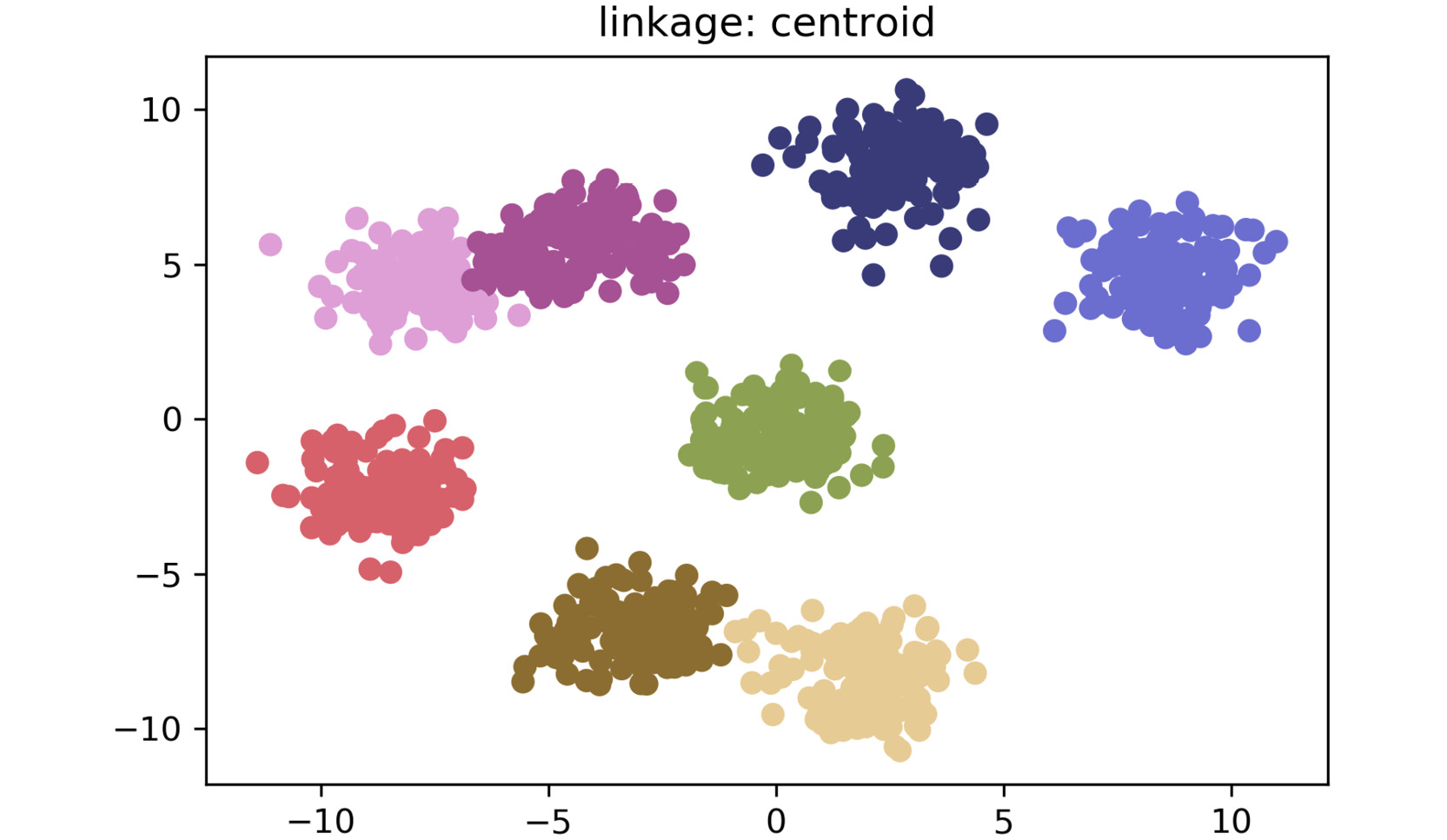
plt.show()The plot for centroid linkage is as follows:
Figure 2.15: A scatter plot for centroid linkage method
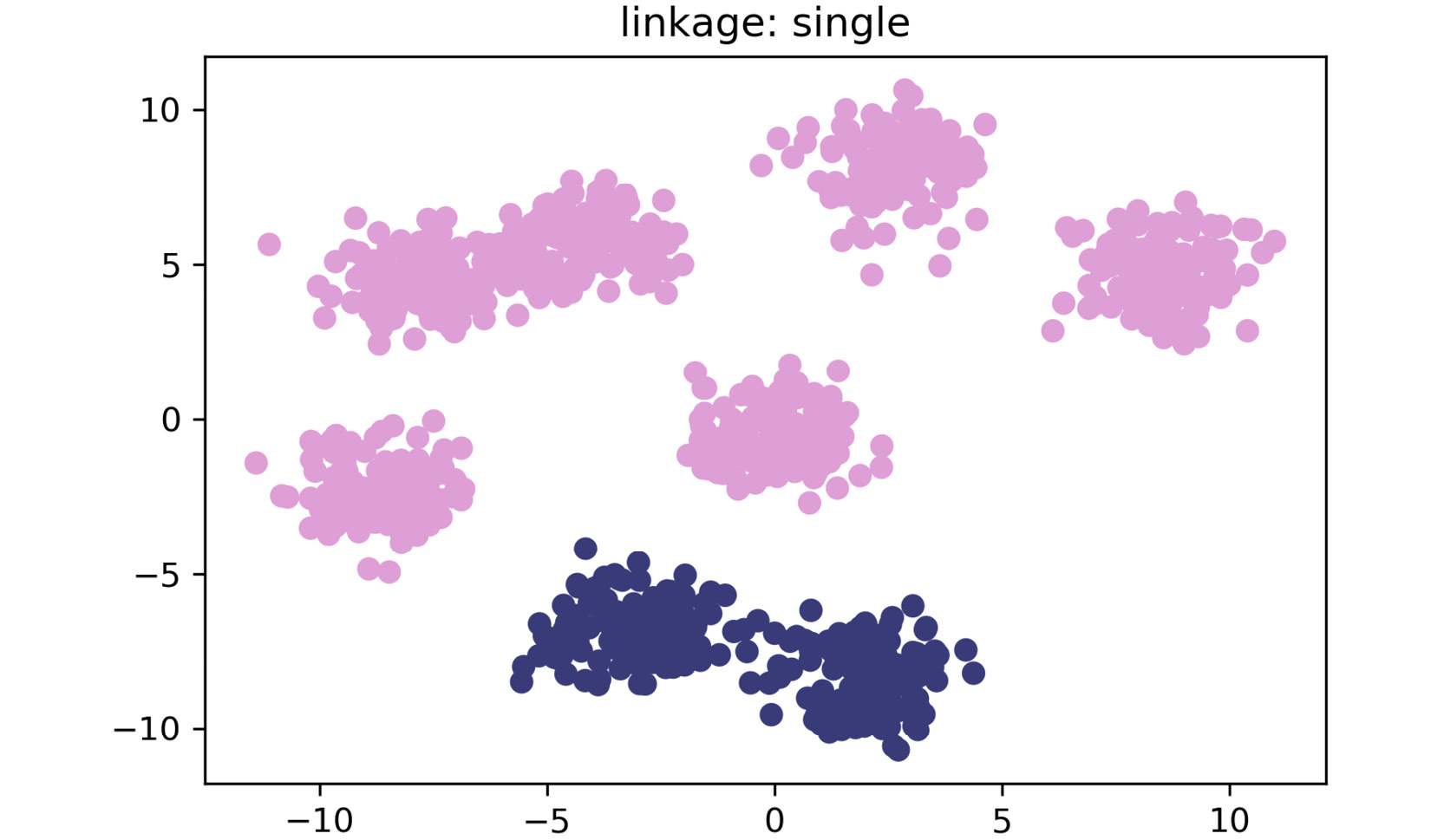
The plot for single linkage is as follows:
Figure 2.16: A scatter plot for single linkage method
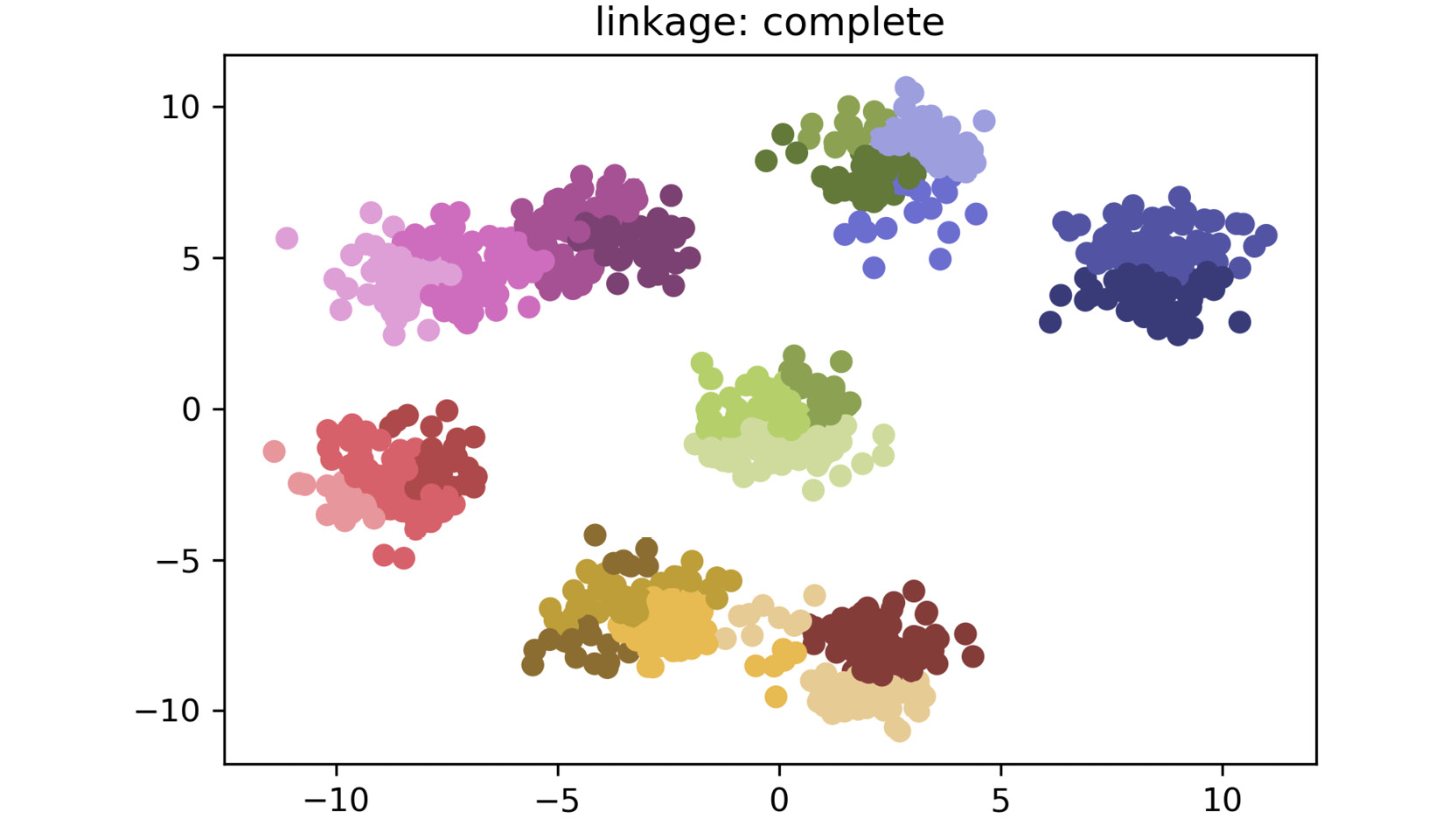
The plot for complete linkage is as follows:
Figure 2.17: A scatter plot for complete linkage method
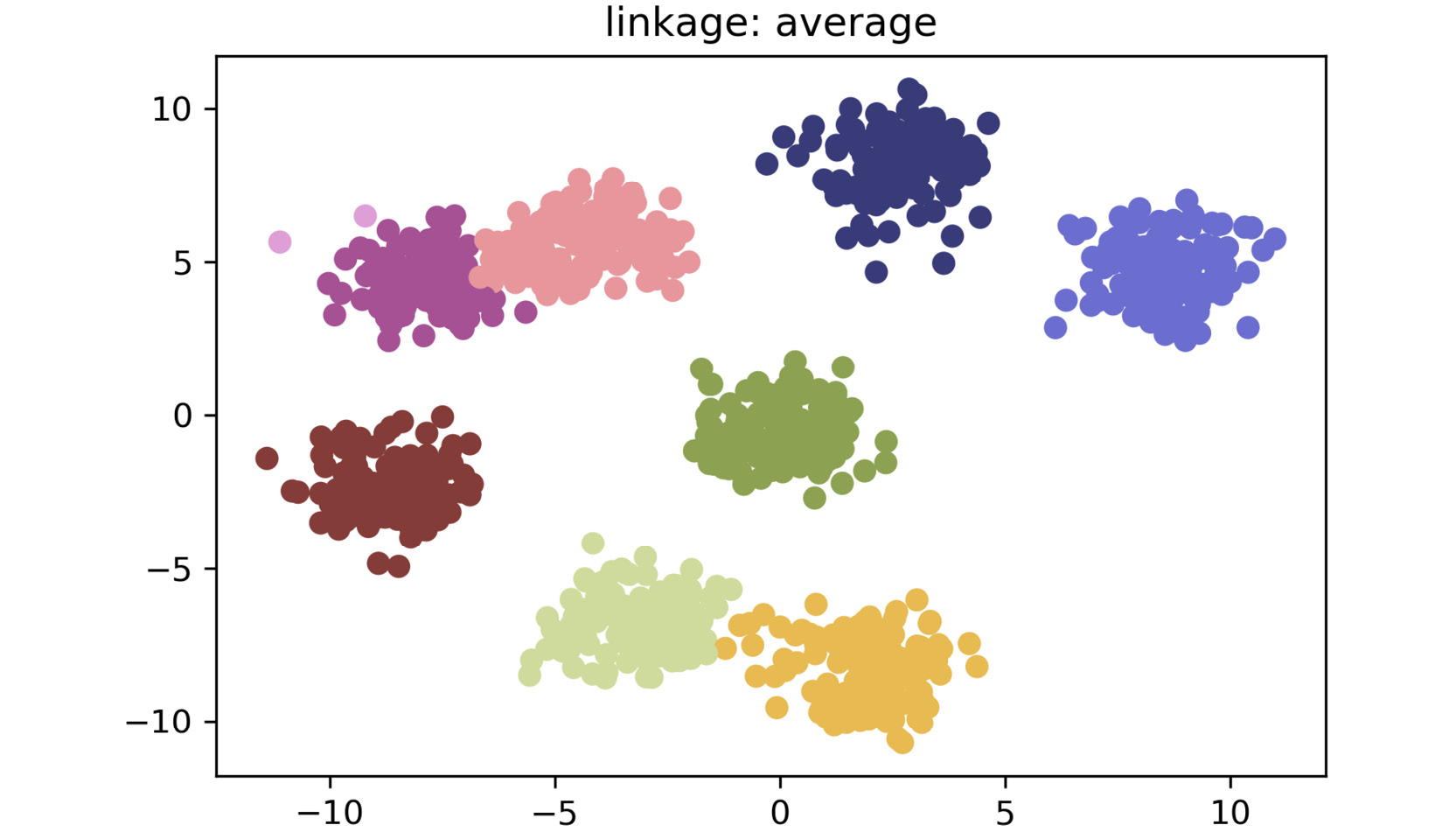
The plot for average linkage is as follows:
Figure 2.18: A scatter plot for average linkage method
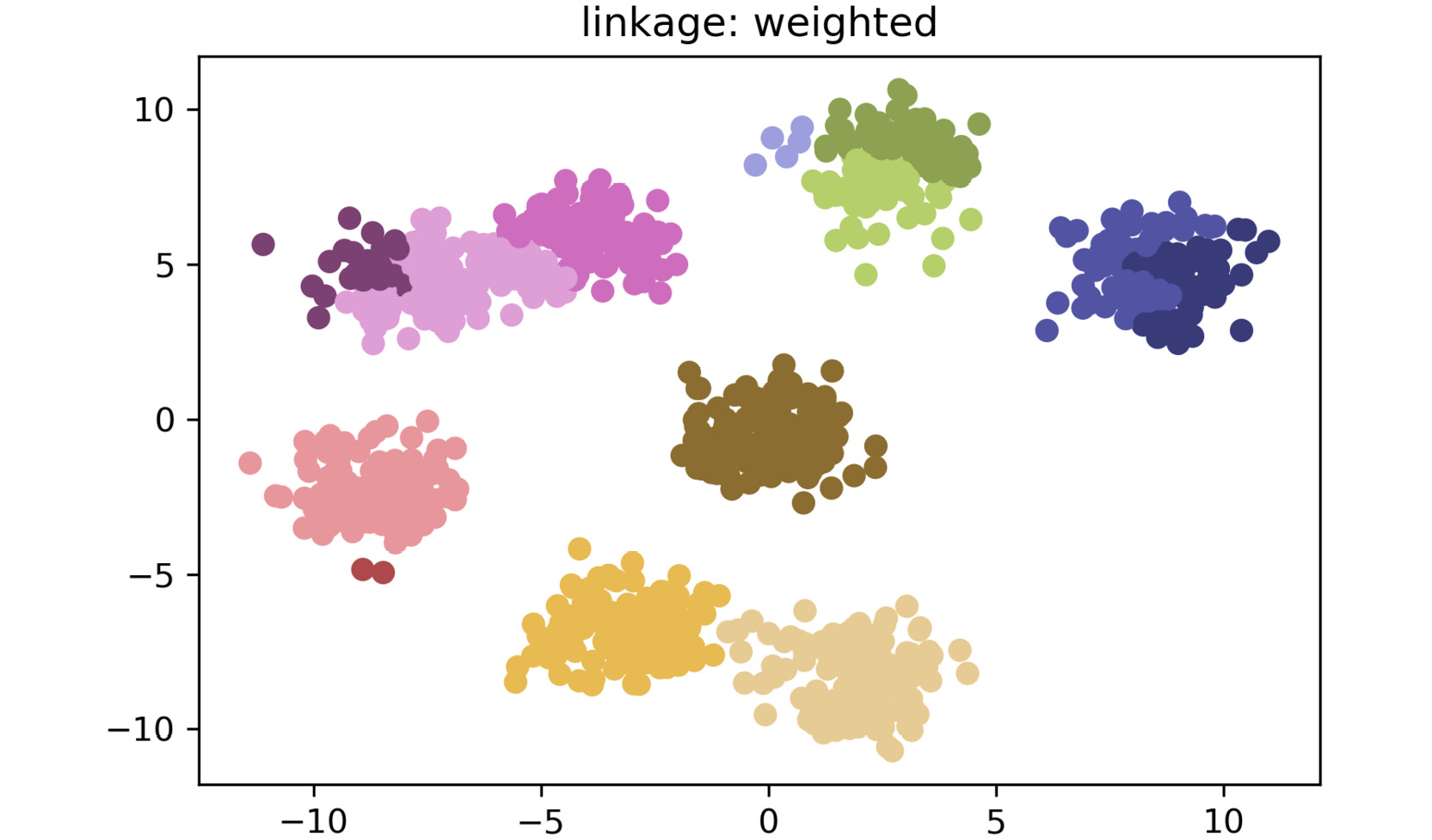
The plot for weighted linkage is as follows:
Figure 2.19: A scatter plot for weighted linkage method
As shown in the preceding plots, by simply changing the linkage criteria, you can dramatically change the efficacy of your clustering. In this dataset, centroid and average linkage work best at finding discrete clusters that make sense. This is clear from the fact that we generated a dataset of eight clusters, and centroid and average linkage are the only ones that show the clusters that are represented using eight different colors. The other linkage types fall short – most noticeably, single linkage. Single linkage falls short because it operates on the assumption that the data is in a thin "chain" format versus the clusters. The other linkage methods are superior due to their assumption that the data is coming in as clustered groups.
Note
To access the source code for this specific section, please refer to https://packt.live/2VWwbEv.
You can also run this example online at https://packt.live/2Zb4zgN.



 Free Chapter
Free Chapter