Let's begin by importing the required packages and preparing the data:
- Let's import the required libraries, functions and classes:
import pandas as pd
import numpy as np
from sklearn.model_selection import train_test_split
from sklearn.impute import MissingIndicator
from feature_engine.missing_data_imputers import AddNaNBinaryImputer
- Let's load the dataset:
data = pd.read_csv('creditApprovalUCI.csv')
- Let's separate the data into train and test sets:
X_train, X_test, y_train, y_test = train_test_split(
data.drop('A16', axis=1), data['A16'], test_size=0.3,
random_state=0)
- Using NumPy, we'll add a missing indicator to the numerical and categorical variables in a loop:
for var in ['A1', 'A3', 'A4', 'A5', 'A6', 'A7', 'A8']:
X_train[var + '_NA'] = np.where(X_train[var].isnull(), 1, 0)
X_test[var + '_NA'] = np.where(X_test[var].isnull(), 1, 0)
Note how we name the new missing indicators using the original variable name, plus _NA.
- Let's inspect the result of the preceding code block:
X_train.head()
We can see the newly added variables at the end of the dataframe:
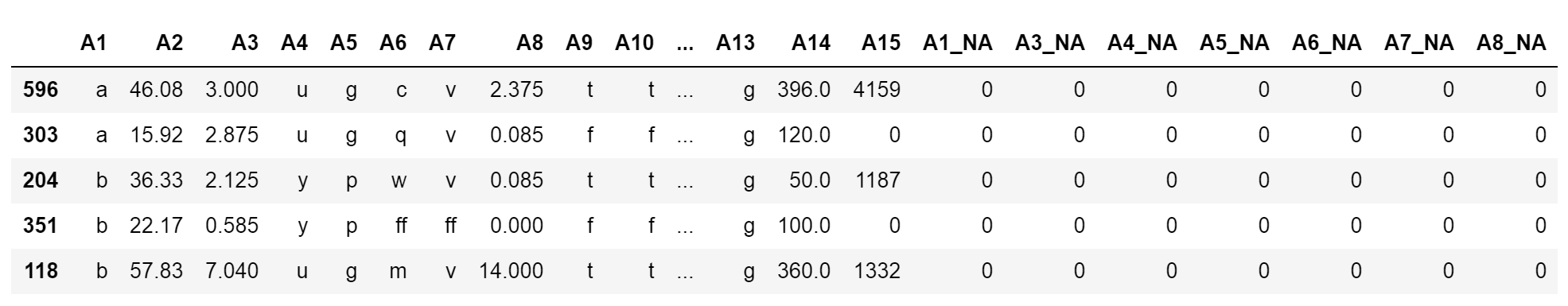
The mean of the new variables and the percentage of missing values in the original variables should be the same, which you can corroborate by executing X_train['A3'].isnull().mean(), X_train['A3_NA'].mean().
Now, let's add missing indicators using Feature-engine instead. First, we need to load and divide the data, just like we did in step 2 and step 3 of this recipe.
- Next, let's set up a transformer that will add binary indicators to all the variables in the dataset using AddNaNBinaryImputer() from Feature-engine:
imputer = AddNaNBinaryImputer()
We can specify the variables which should have missing indicators by passing the variable names in a list: imputer = AddNaNBinaryImputer(variables = ['A2', 'A3']). Alternatively, the imputer will add indicators to all the variables.
- Let's fit AddNaNBinaryImputer() to the train set:
imputer.fit(X_train)
- Finally, let's add the missing indicators:
We can inspect the result using X_train.head(); it should be similar to the output of step 5 in this recipe.
We can also add missing indicators using scikit-learn's MissingIndicator() class. To do this, we need to load and divide the dataset, just like we did in step 2 and step 3.
- Next, we'll set up a MissingIndicator(). Here, we will add indicators only to variables with missing data:
indicator = MissingIndicator(features='missing-only')
- Let's fit the transformer so that it finds the variables with missing data in the train set:
indicator.fit(X_train)
Now, we can concatenate the missing indicators that were created by MissingIndicator() to the train set.
- First, let's create a column name for each of the new missing indicators with a list comprehension:
indicator_cols = [c+'_NA' for c in X_train.columns[indicator.features_]]
The features_ attribute contains the indices of the features for which missing indicators will be added. If we pass these indices to the train set column array, we can get the variable names.
- Next, let's concatenate the original train set with the missing indicators, which we obtain using the transform method:
X_train = pd.concat([
X_train.reset_index(),
pd.DataFrame(indicator.transform(X_train),
columns = indicator_cols)], axis=1)
Scikit-learn transformers return NumPy arrays, so to concatenate them into a dataframe, we must cast it as a dataframe using pandas DataFrame().
The result of the preceding code block should contain the original variables, plus the indicators.



 Free Chapter
Free Chapter

